First of all, click on the "New Job" button:
A screen to define your job parameters will be opened. Define there the UniProt accession
of your target protein (mandatory).
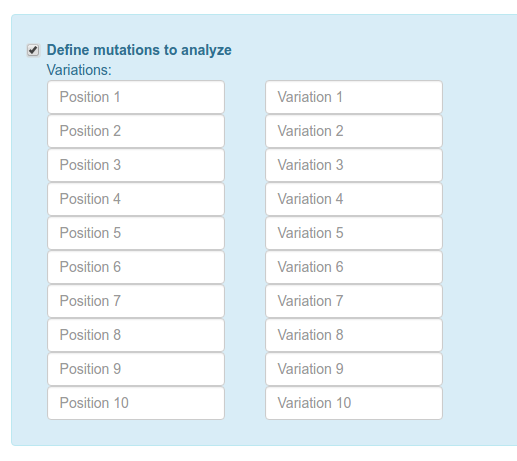
Then if you want to search mutations in public databases(Be aware that widely studied proteins have hundred of reported
mutations and some of the jobs very slow) and/or define specific mutations to be analyzed (up to 10 per job).


Finally, write your mail to be informed about your job status, and send to run!